|
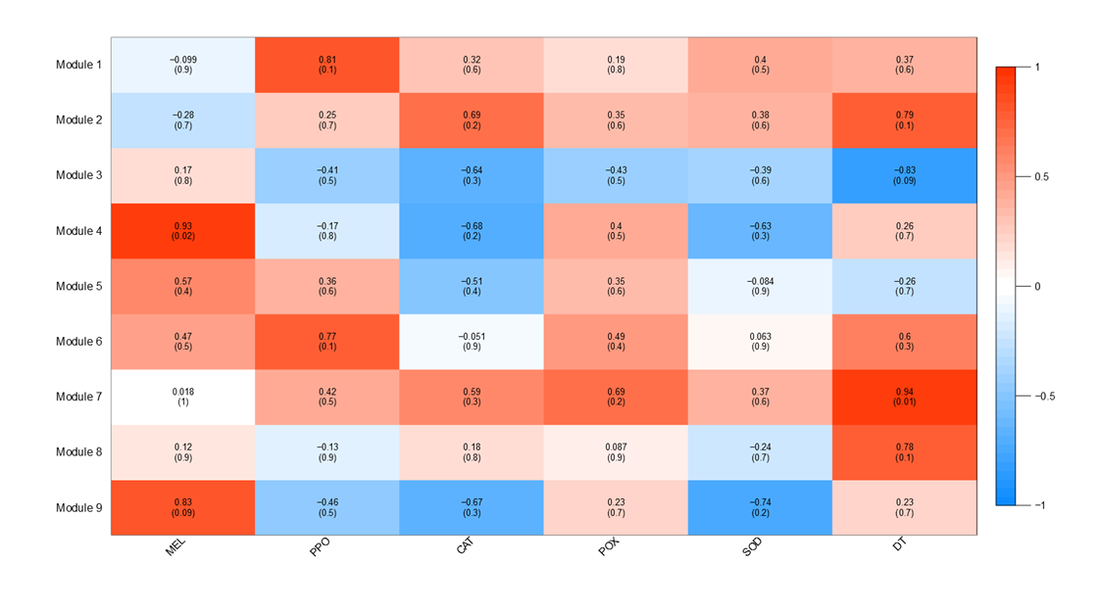
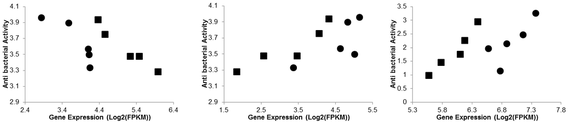
Just a few weeks ago, my newest publication was accepted for publication in Developmental and Comparative Immunology. If you haven't read it yet, you can check it out here or go to my publication page and download the pdf! To summarize the paper: we used novel correlative and network based analyses to connect gene expression to protein data. It may seem straight forward enough, but working on such things in a non-model organism was certainly a challenge! It took a lot of creativity, but in the end we were able to achieve our goal of identifying new transcripts which change in expression during immune challenge and were correlated to immune protein activity! Now that the paper's out, I thought I'd devote a blog post to detailing the journey we took to get to that final goal. When I first started my graduate career, my main goal was to go beyond the, at that time, standard in transcriptional studies of non-model organisms. I was hoping to move beyond lists of deferentially expressed genes. Namely I wanted to employ more "genome to phenome" type techniques, so to speak, in order to use transcriptomics to really get at ecological questions. This new paper was my first attempt at doing just that. At SICB in 2014, I first learned of a new R package, Weighted Gene Correlation Network Analysis, which not only groups genes based on patterns of expression, but also allows for the correlation of those groups of genes to quantitative traits. This was just the kind of tool I was looking for, so naturally I wanted to try it out. Our post-doc at the time had the same idea and was kind enough to collaborate with me on this project using a great set of data from an immune challenge experiment that had previously been conducted. In the process of analyzing this dataset using WGCNA, I learned a lot about statistics, and the short comings of working in a non-model organism for which there are few gene annotations. Essentially, while WGCNA worked great at grouping genes and correlating them to our protein assay results, the lack of gene annotations made it almost impossible to draw any significant conclusions using that software alone. But, as you can see from my supplementary tombstone plots (Figure 1), there were lots of groups of genes which were significantly correlated to types of immunity! The problem was we just couldn't characterize these groups well. Figure 1: Supplementary figure displaying correlations between modules and different measures of immunity. When we realized that WGCNA just couldn't quite get us to where we wanted to go, I once again had to get creative with my analysis techniques. This time I tried something I hadn't seen done much before: directly correlating gene expression to immune activity. WGCNA became a filter: all the genes in significant modules were screened for individual correlations to a measure of immune activity. This single gene-protein correlation technique allowed us to identify genes which not only changed during immune challenge, but also had correlations to functional measures of immunity (Figure 2). Using this technique we were able to add confidence to the identification of potential candidate immune genes beyond analysis of blast annotation results. In addition, using this combination of differential expression and correlative techniques, we were able identify genes which were down-regulated post immune challenge but were annotated as positive regulators of immunity and correlated positively to functional measures of immunity. While we still don't fully understand these patterns, its possible that disregulation of sorts of these immune genes may be contributing to differential susceptibility of corals to disease. So, while the journey may have been long, and we may have had to get creative at times, I believe this new publication really does make a strong contribution to the field of marine invertebrate immunology. Moral of the story: when using transcriptional techniques to study non-model organisms, don't be afraid to get creative and travel off the beaten path. You never know what you might find! Figure 2: correlations between immune genes (LRR domain protein, DMBT1, HSP70) and measures of immunity (antibacterial activity for the first two, propenoloxidase for the third)
0 Comments
|
AuthorLauren Fuess is a postdoctoral researcher at the University of Connecticut working with Dr. Daniel Bolnick. Formerly she was a PhD student at the University of Texas at Arlington with Dr. Laura Mydlarz. The opinions expressed here are solely her own. Archives
September 2019
Categories |
 RSS Feed
RSS Feed
